During dorsal closure (DC), lateral epidermal sheets advance to close a dorsal opening that is temporarily filled with a squamous layer of amnioserosa cells (see Kiehart et al, Ann. Rev. Cell Dev. Biol. 2017 33:169-202). How does this happen? DC is a dynamic process that is robust and remarkably resilient – no single force that is involved in driving closure is absolutely required. Many intriguing questions in our understanding of this complex cell sheet movement remain, including:
- How are forces from the motor protein nonmuscle myosin 2 regulated and coordinated?
- How are collective dynamics dependent on the constraints generated by the mechanical properties of surrounding tissues and extracellular matrix?
- In the context of global developmental patterning and signaling events, how do forces produced by the actomyosin cytoskeleton result in individual and collective cell movements and shape changes?
We use a multidisciplinary approach to investigate the mechanisms of DC and morphogenesis across phylogeny.
As one example of our forward genetics approach, we are generating a more complete ‘parts list’ of genes that contribute to closure. To identify new “DC genes” (i.e., genes that when deleted, disrupt closure), we have expanded our genetic and biophysical studies of DC with a forward genetics, live-imaging, pilot screen. This screen utilizes video analysis of DC in embryos that are homozygous for genetic deficiencies (Dfs), which remove contiguous portions of the genome. This allows us to evaluate the contribution of all of the genes deleted by the Df after just two crosses (designed to introgress the Df into an “imaging background”) and to uncover roles for genes in that region during DC, whether they have subtle or severe effects on closure. We have screened 5,420 genes (>92.6%) of the 5,854 genes on the 2nd chromosome for DC defects and categorized these Dfs based on the tissues or processes effected (summarized in the pie chart below). We use overlapping Dfs, existing mutations or generate new CRISPR-based knockouts to narrow down the gene(s) responsible for the most interesting and informative, observed DC defects. In some cases, the genetic cause for complex phenotypes can be sorted into simpler phenotypes with the use of sub-Dfs that collectively span the parent Df initially used in the screen. To date, we have identified 7 DC genes or “pre-DC” genes on the 2nd chromosome: even-skipped, short stop, tumbleweed, three rows, jelly belly, odd-skipped, sloppy-paired 1, pimples, and paired. Based on our 2nd chromosome pilot screen, we extrapolate to conclude a Df screen of all 4 chromosomes in Drosophila will yield approximately 165 new DC genes!

Pie chart summary of Df screen phenotypes for chromosome 2.
(Mortensen et al., 2018. G3. 8(7), 2361-2387) and
Fogerson et al., 2020. G3. Accepted for publication.
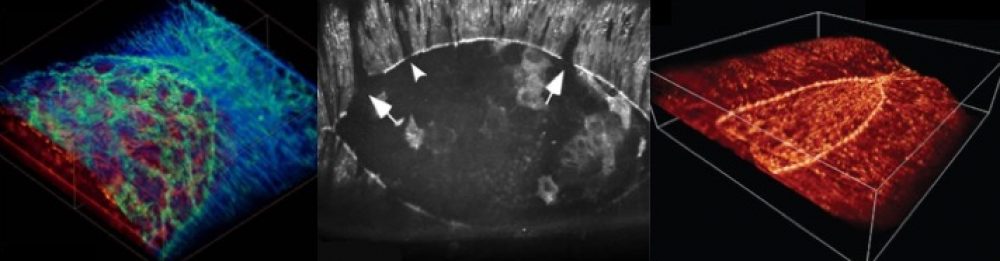
Once we know the players involved, to understand their roles, we assess how mutations in those players affect the machinery of development. To this end, we apply our biophysical approaches including super-resolution microscopy, laser microsurgery and mechanical probing to measure motions, forces and mechanical properties of the tissues involved in DC.
A new project in the lab provides an example of a directed molecular approach and focuses on motor function by nonmuscle myosin 2: the fly uses 3 different isoforms of myosin 2, generated through the differential inclusion of exons in Loop 1 of the myosin 2 motor domain. We are investigating where the spliceforms are expressed and how they function in living flies. We are also using fast time course kinetics and in vitro motility assays to assess the extent to which the different versions of Loop 1 affect myosin motor function.